Remedial strategy for chlorinated solvent sites typically requires a combination of technologies to achieve site-specific goals for closure. Additionally, a majority of these sites will have monitored natural attenuation (MNA) as a remedy component and need to be evaluated for the lines of evidence required to establish the viability of MNA. Based on Haley & Aldrich research and findings, presented at the AEHS East Conference in October 2019, microbiological tools (MBTs), such as quantitative polymerase chain reaction (qPCR) and 16S ribosomal gene sequencing, are essential components for remediation practitioners to help with the strategy and design of in-situ remediation as well as the MNA path forward. MBTs have been used in the environmental industry for years and, as analytical methods have continued to be developed, the variety of MBTs available has risen while the costs have decreased significantly from when they were first introduced. The availability and relatively inexpensive cost of MBTs have made them indispensable in formulating remediation strategies for chlorinated solvent sites.
MBTs provide a snapshot of the bacterial populations present, which helps to clarify the natural attenuation processes occurring in the subsurface and allows for more robust and comprehensive knowledge of the site when combined with other analytical and site data. Remedial technology selection can be informed by the use of MBTs, such as whether additional electron donors and/or nutrients that may be lacking should be added, or if an anaerobic or reductive approach is more appropriate at a site where an in-situ chemical oxidation event could potentially disrupt a flourishing anaerobic population. A more complete and accurate picture of the subsurface can be gained when using MBTs, which allow for a better evaluation of potential remedial actions and natural attenuation. MBTs provide the ability to:
- design the optimal and most effective in-situ remediation program that aligns with the site strategy,
- obtain a snapshot of biological populations to make decisions on remedial progress,
- predict when to amend the subsurface conditions to enhance the growth of the “right bugs” following aggressive remedial technology applications,
- identify additional bacterial populations that may be useful in the biodegradation of the site contaminants via direct or indirect pathways, and
- determine whether additional amendments are necessary, or an MNA remedy could be pursued.
Microbiological tool selection
We have been utilizing the MBTs for remedial decision making and the evaluation of natural attenuation. The three methods listed below are readily and commercially available by major laboratories and provide different levels of data:
- One DNA-based technique used to detect and quantify specific microorganisms or functional genes is qPCR. Examples include quantifying Dehalococcoides mccartyi (DHC) at a chlorinated solvent site.
- Another tool, 16S ribosomal gene sequencing, utilizes 16S ribosomal RNA to provides a broad snapshot of biological presence in the sample and pinpoints species-specific signature sequences to identify bacteria present in the environmental sample.
- Shotgun metagenomic gene sequencing is the genomic analysis of microbial DNA from environmental communities. It is used to examine thousands of organisms in parallel and comprehensively sample all genes, which can allow for insights into the subsurface’s biodiversity and function and provide information on all of the DNA found in the sample.
The use of MBTs when coupled with additional analytical data can provide an extensive amount of information including a snapshot of the current population of bacteria in the subsurface. MBTs such as shotgun metagenomics can provide both an overview of the bacterial population present in the subsurface as well as the quantitative concentrations of the specific bacteria present. In addition, these techniques can provide insight into the functional genes present in the bacteria, which can allow for further information on the ability of the bacteria to degrade the contamination present at a site. These tools enable better decisions to be made throughout the remediation life-cycle. They can inform an optimal approach and design, determine whether additional remedial activities are necessary, ascertain if bioaugmentation (the addition of cultured microorganisms into the subsurface to biodegrade soil and groundwater contaminants) is required or may be beneficial, and evaluate natural attenuation pathways for site closure. The case studies below illustrate the benefit of utilizing MBTs.
Case study: Public university site, New Jersey
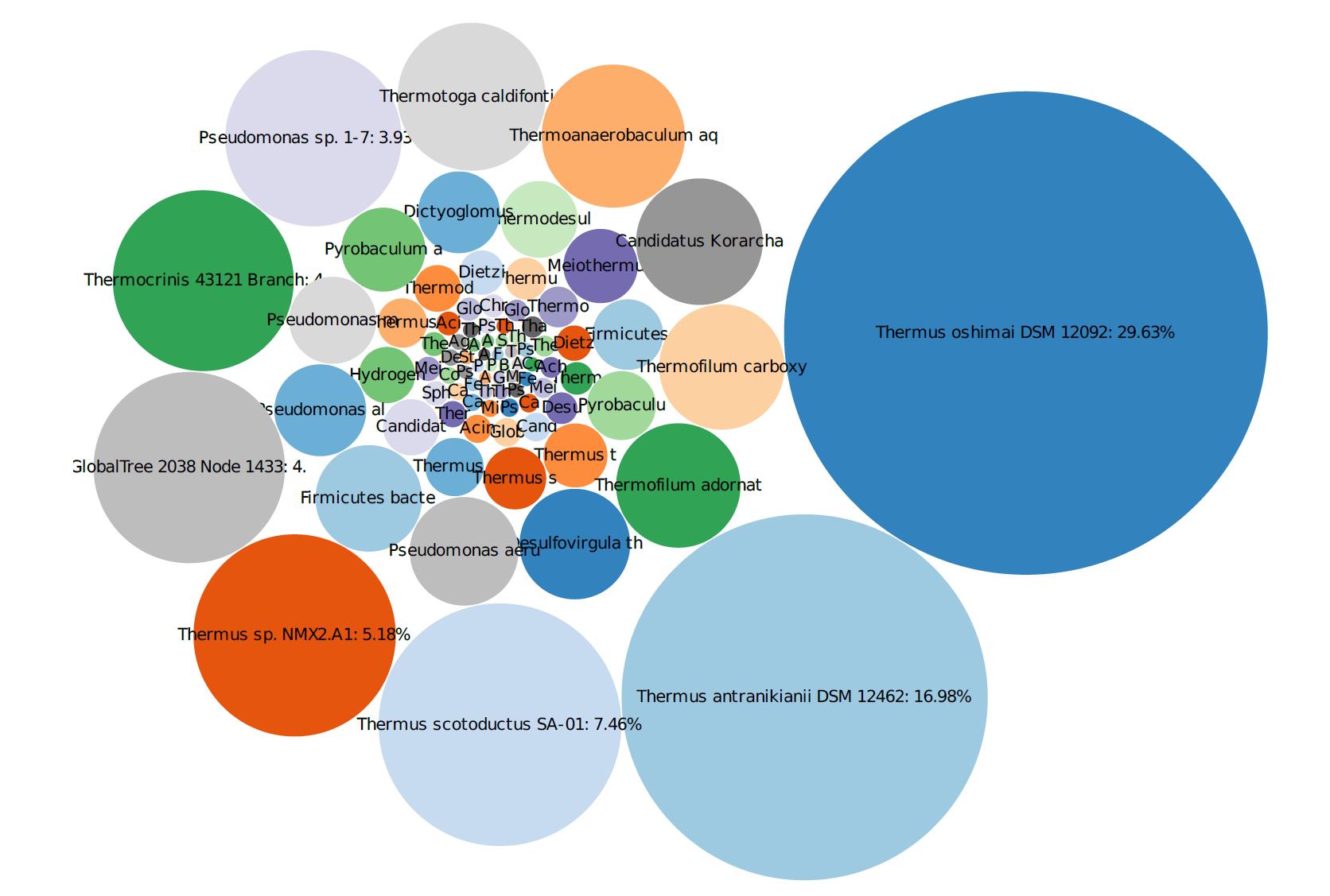
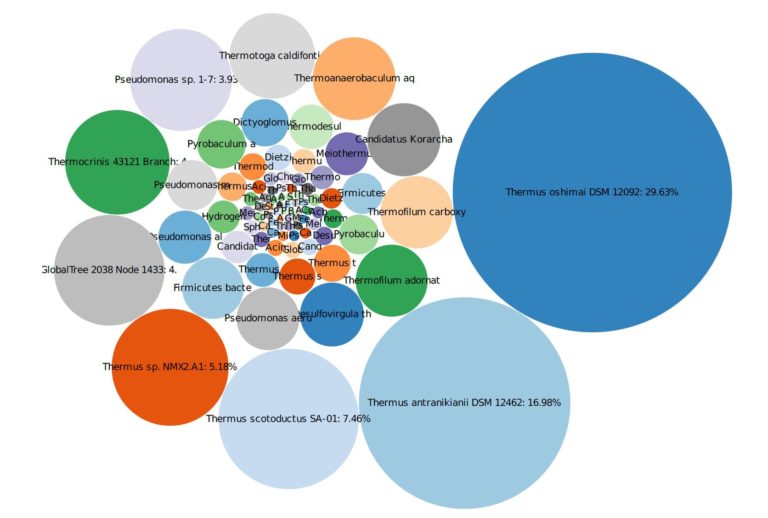
Following an electrical resistance heating (ERH) remedial program, MBTs together with VOC and geochemical analyses were used to determine the need and timing of an additional polishing step following the aggressive source reduction. The site was contaminated with tetrachloroethene (PCE) and carbon tetrachloride (CT), and their corresponding daughter products. The data utilizing MBTs, specifically qPCR, 16S gene sequencing, and shotgun metagenomics, was collected several times post-ERH to determine the biological population and diversity as the site was cooling. The results indicated that the subsurface was initially too warm within the treatment area at one-year post-ERH as there were high concentrations of thermophiles present, though the fringes of the treatment area showed elevated concentrations of DHC and other known dechlorinating bacteria. The MBT data provided the information to plan and time the remedial event optimally and decide on bioaugmentation that the VOC and geochemical data would not have provided. In addition, using metagenomic testing allowed for an overview of the biodiversity that would have been missed if just qPCR were utilized.
Case study: Confidential site, Arizona
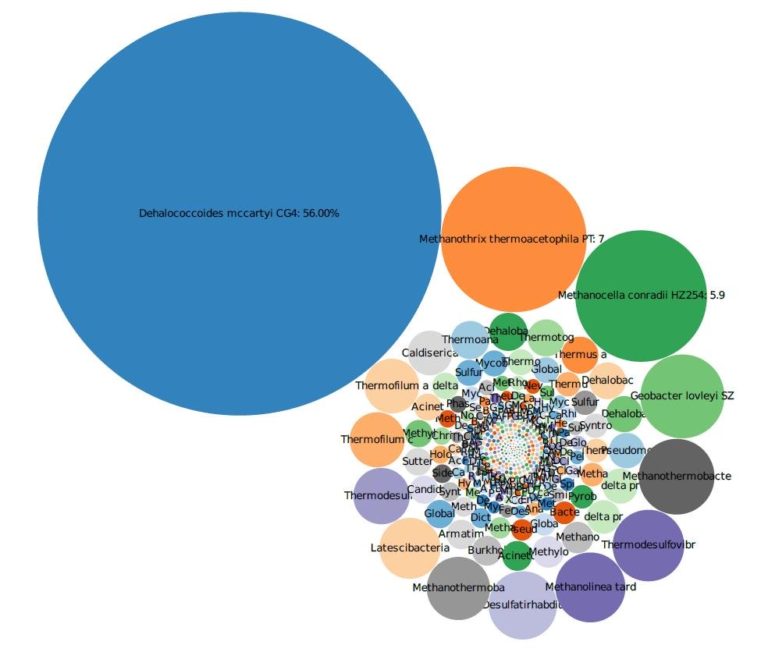
The site was contaminated with low levels of trichloroethene (TCE) (<20 ug/L) and its respective daughter products. To evaluate MNA, we used shotgun metagenomic and 16S gene sequencing to determine if MNA was a viable pathway for site closure. The results show that low concentrations of known TCE degraders such as DHC were present; however, results also showed that elevated concentrations of bacteria capable of co-metabolic degradation of the contamination were also present. The presence of these bacteria-which can degrade the contamination present independent of the concentration of the contaminants-provided the tertiary line of evidence necessary to obtain regulatory approval for the MNA program.
Case study: Public university site, New Jersey
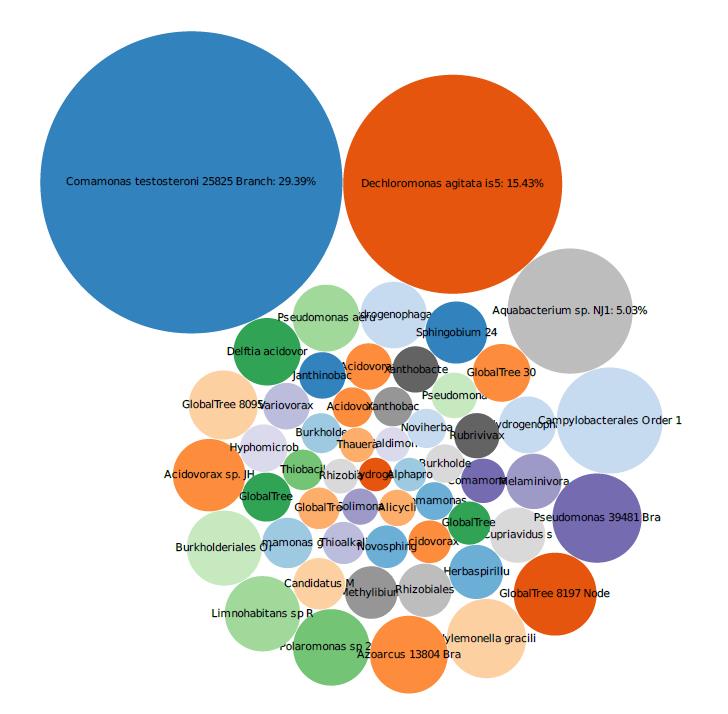
The site was contaminated with elevated concentrations of CT within bedrock and had been treated previously with an alkaline activated sodium persulfate oxidation, which resulted in a reduction of source mass concentrations. Natural attenuation was evaluated by reviewing the VOC, geochemical, and biological data via shotgun metagenomic and biomass data collected at the site. The metagenomic data indicated that low populations of bacteria capable of biodegradation of CT were present in the subsurface and that lines of evidence for MNA were not strong. This suggested that additional remedial measures were necessary. The data from MBTs provided the strategy and design for the path forward, which included injection electron donors and bioaugmentation cultures at the site in August 2019.
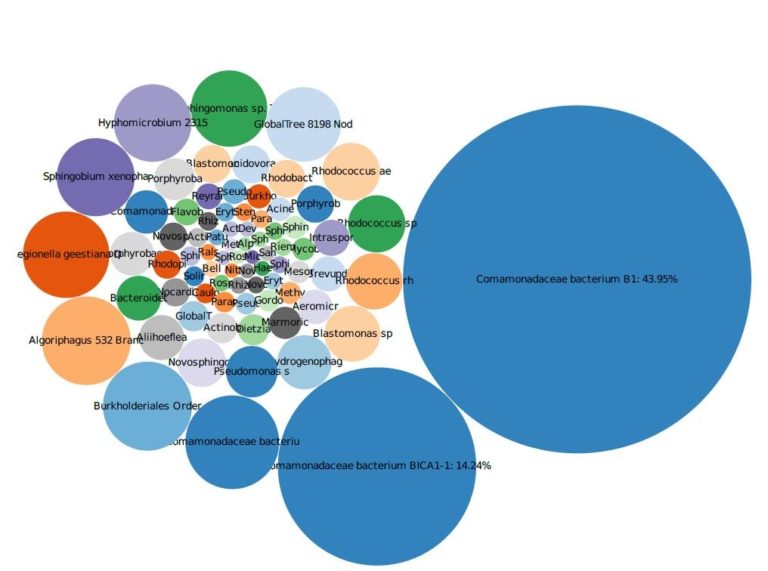
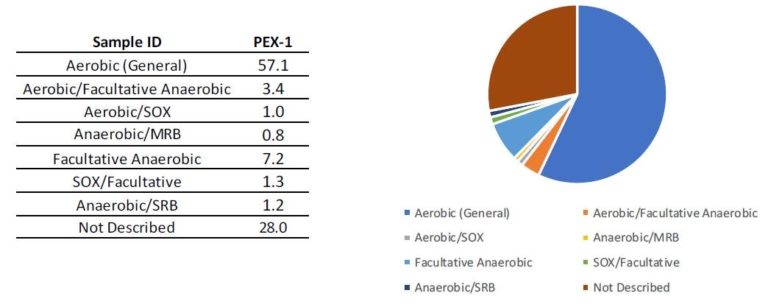
Case study: Confidential site, New Jersey
The site was contaminated with chlorinated solvents and was evaluated for natural attenuation and the need for additional remedial activities. A qPCR array was utilized to evaluate the bacteria populations, in conjunction with contaminant and geochemical data. The results of the analysis indicated that low concentrations of bacteria capable of biodegradation were present but at concentrations that needed to be stimulated. We used this data to design a biodegradation remedial event implemented in the summer of 2019. Additional MBTs will be used in the future to evaluate the remedy and determine when additional amendment and bioaugmentation cultures should be injected into the subsurface.
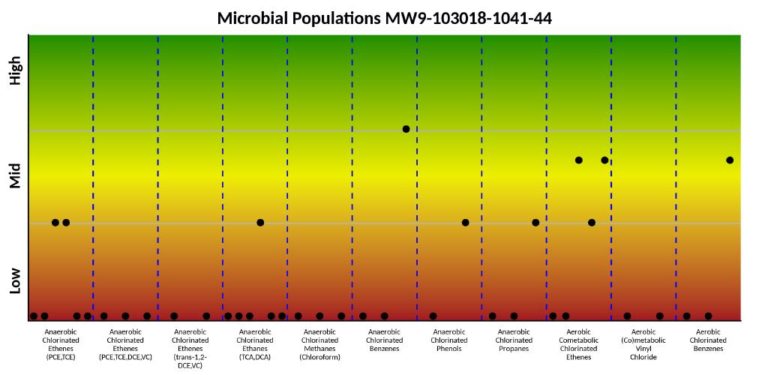
Conclusions
MBTs are a crucial and important component of a remediation practitioner’s toolbox to help determine what is occurring in the subsurface and inform remedial strategies and decisions. MBTs can help fill critical data gaps present on remediation sites and enable a more complete understanding of degradation pathways in the subsurface. The use of one or more of tools such as qPCR, 16S ribosomal gene sequencing, and shotgun metagenomic gene sequencing can be a relatively inexpensive way to determine how in-situ bioremediation fits into the overall remedial and closure strategy. Additionally, they can evaluate conditions in the subsurface that are not conducive for the desired bacterial populations (e.g. the subsurface is too warm following ERH) as well as assess the overall microbial population and whether the lack of the “right” bacteria in optimal quantities is truly impeding contaminant degradation.
Contact Haley & Aldrich’s Program Manager, Sunila Gupta, to discover how MBTs are an important tool to help inform remedial strategies and decisions.
Published: 10/22/2019
- Aerospace
- Industrial and manufacturing
- Automotive
- Consumer products
- Food and beverage
- Manufacturing
- Contaminated site management
- Remediation
Authors

Program Manager

Senior Technical Specialist